
Contact Information
Name: Kimberly Mulligan
Title: Professor
Email: kimberly.mulligan@csus.edu
Mailing Address: Sacramento State 6000 J Street Sacramento, CA 95819-6077
Office Hours: Please email for current office hours information
Courses that I Teach
- BIO 294A: Seminar in Molecular and Cellular Biology
- BIO 220: Introduction to Scientific Inquiry
- BIO 227: Development and Regenerative Medicine
- BIO 127: Developmental Biology
- BIO 121: Molecular Cell Biology
Lab Homepage
https://www.mulliganlab.com/Publications
Welch C and Mulligan K (2022) Evaluating learning and memory in Drosophila melanogaster to study the neurodevelopmental impacts of toxicants. Current protocols, 2(10), e576. https://doi.org/10.1002/cpz1.576
Welch, C.; Mulligan, K.(2022) Does Bisphenol A Confer Risk of Neurodevelopmental Disorders? What We’ve Learned from Developmental Neurotoxicity Studies in Animal Models. Invited Review. International Journal of Molecular Sciences, 23(5), 2894. https://doi.org/10.3390/ijms23052894
Mulligan K and Cheyette B (2016) Introduction to Wnt signaling, Chapter for Inborn Errors of Development, 3rd Edition, Oxford University Press
Martin P, Stanley R, Ross A, Freitas A, Moyer C, Brumback A, Iafrati J, Stapornwongkul K, Dominguez S, Kivimae S, Mulligan K, Pirooznia M, McCombie W, Potash J, Zandi P, Purcell S, Sanders S, Zuo Y, Sohal V, Cheyette B (2016) DIXDC1 contributes to psychiatric susceptibility by regulating dendritic spine and glutamatergic synapse density via GSK3 and Wnt/β-catenin signaling Mol Psychiatry. Oct 18. doi: 10.1038
Mulligan K and Cheyette B (2012) Wnt signaling in vertebrate neural development and function Review. J NeuroImmune Pharmacol. Dec; 7(4) 774-87
Mulligan K, Fuerer C, Ching W, Willert K, Fish M, Nusse R (2012) Secreted-Wingless interacting molecule (Swim) promotes long-range signaling by maintaining Wingless solubility Proc Natl Acad Sci USA. Jan10;109(2):370-7
Nusse R, Fuerer C, Ching W, Harnish K*, Logan C, Zeng A, ten Berge D, Kalani Y. (2008) Wnt signaling and stem cell control Cold Spring Harb Symp Quant Biol. Nov (73) 59-66. Review
Johnson ML, Harnish K*, Nusse R, Van Hul W (2004) LRP5 and Wnt signaling: a union made for bone. J Bone Mineral Research. Nov;19(11):1749-57. Review
Research Projects/Interests
My lab is interested in understanding how genes and non-heritable environmental factors may converge to cause neurodevelopmental disorders (NDDs). Many NDDs are complex, multifactorial disorders caused when multiple genes and environmental factors converge to affect the normal neurodevelopmental program during prenatal brain development. The identification of environmental factors that confer risk of NDDs and analysis of their molecular mechanisms could provide measures to prevent or decrease the severity of NDDs.
In our lab, we use the common fruit fly (Drosophila melanogaster) to identify and examine candidate environmental chemicals to determine if they can impair neural development in flies that have genetic mutations associated with NDDs. It is estimated that 75% of the genes associated with human disease are conserved in Drosophila, and they have already served as a model organism for the study of genes implicated in NDDs in humans. NDD susceptibility genes in flies can cause measurable social and behavioral deficits that can be examined in a lab setting. Fruit flies are also incredibly easy to manipulate genetically, making them ideal for the study of multiple genetic mutations in the same organism. There are a number of well-defined behavioral assays, one of the most prominent being the courtship assay. (Flies have specific, stereotyped behaviors they perform during courtship and alteration in these behaviors is often indicative of a change in neural circuitry.) If behavioral changes are detected in adult flies, brains and neurons can be examined to elucidate the mechanism by which a particular chemical alters brain development. Finally, fruit flies have a generation time from egg to adult of 9 days, so meaningful data can be generated relatively quickly; this is critical given the urgent need to understand the causes of NDDs.
For more information, please visit my lab homepage.
Article About My Teaching Strategies
https://www.coursehero.com/faculty-club/classroom-tips/kimberly-mulligan/Other Information
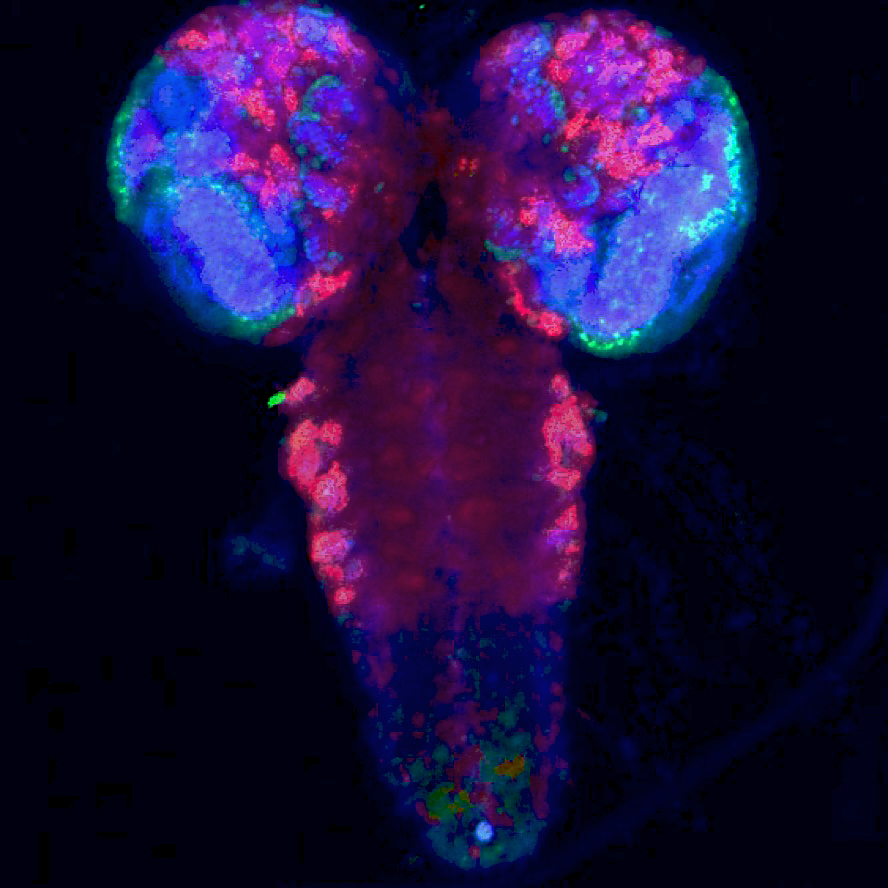
Full view of the fruit fly larval brain.
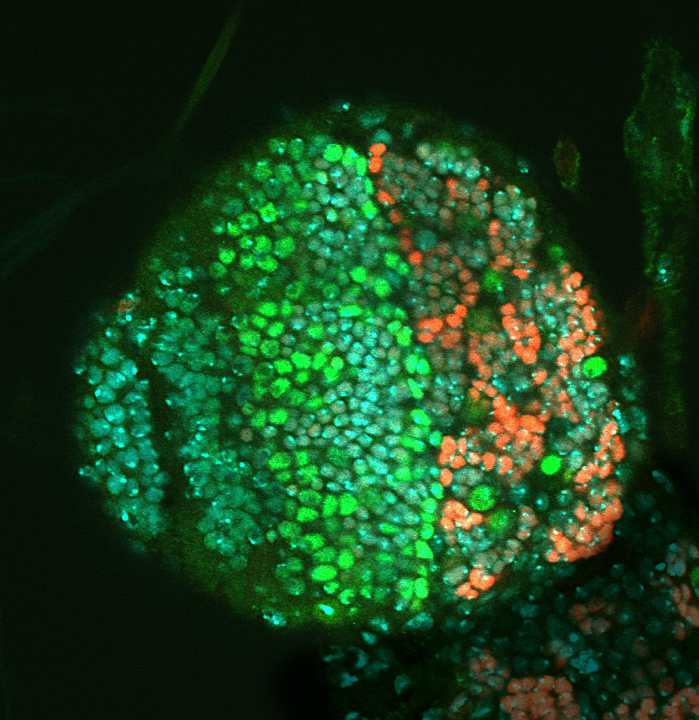
A lobe of the fruit fly larval brain with fluorescently marked neural stem cells.